Automated tissue image analysis 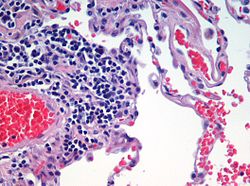 Automated tissue image analysis or histopathology image analysis (HIMA) is a process by which computer-controlled automatic test equipment is used to evaluate tissue samples, using computations to derive quantitative measurements from an image to avoid subjective errors. In a typical application, automated tissue image analysis could be used to measure the aggregate activity of cancer cells in a biopsy of a cancerous tumor taken from a patient. In breast cancer patients, for example, automated tissue image analysis may be used to test for high levels of proteins known to be present in more aggressive forms of breast cancers. ApplicationsAutomated tissue imaging analysis can significantly reduce uncertainty in characterizing tumors compared to evaluations done by histologists,[1] or improve the prediction rate of recurrence of some cancers.[2][3] As it is a digital system, suitable for networking, it also facilitates cooperative efforts between distant sites.[4] Systems for automatically analyzing tissue samples also reduce costs and save time.[1] High-performance CCD cameras are used for acquiring the digital images. Coupled with advanced widefield microscopes and various algorithms for image restoration, this approach can provide better results than confocal techniques at comparable speeds and lower costs.[5] ProcessesThe United States Food and Drug Administration classifies these systems as medical devices, under the general instrumentation category of automatic test equipment.[6] ATIS have seven basic processes (sample preparation, image acquisition, image analysis, results reporting, data storage, network communication, and self-system diagnostics) and realization of these functions highly accurate hardware and well-integrated, complex, and expensive software.[7] PreparationSpecimen preparation is critical for evaluating the tumor in the automated system. In the first part of the preparation process the biopsied tissue is cut to an appropriate size (typically 4 mm), fixed in buffered formalin, dehydrated in ethanol-xylene, embedded in paraffin, thin sectioned typically to 4 um slices, then mounted onto at least two barcoded slides (a control and a test). Next the paraffin is removed from the tissue, the tissue is rehydrated, then stained. Any inconsistency in these procedures from case to case may result in uncertainties in the outcome of the analysis. These potential and irreducible inconsistencies in analysis results motivated the development of Automated Tissue Image Systems. AcquisitionDigital micrographs are acquired of the stained specimen on the glass slide. The images are taken by a set of charge-coupled devices (CCD).[8] AnalysisImage analysis involves complex computer algorithms which identify and characterize cellular color, shape, and quantity of the tissue sample using image pattern recognition technology based on vector quantization. Vector representations of objects in the image, as opposed to bitmap representations, have superior zoom-in ability. Once the sample image has been acquired and resident in the computer's random access memory as a large array of 0's and 1's, a programmer knowledgeable in cellular architecture can develop deterministic algorithms applied to the entire memory space to detect cell patterns from previously defined cellular structures and formations known to be significant.[9] See alsoReferences
External links
|