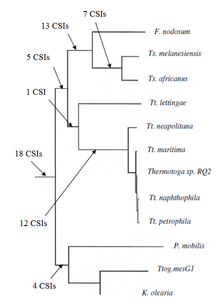
Conserved signature indels
|
Read other articles:

Georg Bleibtreu Georg Bleibtreu (27 Maret 1828 – 16 Oktober 1892) adalah seorang pelukis adegan militer dan sejarah Jerman. Biografi Lahir di Xanten pada 27 Maret 1828), Bleibtreu adalah seorang pelukis, litografer, desainer dan 'graveur sur bois'. Dia adalah anggota Akademi Berlin, menerima dua medali emas untuk karyanya; ia juga menerima medali di Wina pada tahun 1873. Seperti banyak pelukis sejarah abad ke-19, ia belajar di Akademi Düsseldorf antara tahun 1843 dan 1848, da...

Peta laut Peta laut adalah proyeksi[1] bumi atau sebagian muka bumi yang di gambarkan di atas bidang datar dan digunakan untuk berlayar di laut. Peta laut dibuat sedemikian rupa sehingga dapat dipakai untuk merencanakan suatu pelayaran baik di laut, lepas pantai maupun di perairan umum. Peta laut merupakan salah satu alat bantu navigasi untuk keselamatan pelayaran. Di Indonesia yang berhak menerbitkan peta laut adalah Dinas Hidro-Oseanografi TNI AL.[2] Ditinjau dari fisiknya p...

For other uses, see When You're Smiling (disambiguation). SongWhen You're SmilingSheet music, 1928SongPublished1928 by Mills MusicSongwriter(s)Larry Shay, Mark Fisher, Joe Goodwin When You're Smiling is a popular song written by Larry Shay, Mark Fisher and Joe Goodwin. First published in 1928, popular recordings were made by Seger Ellis (1928), Louis Armstrong (1929), and Ted Wallace & His Campus Boys (1930).[1] The lyrics and music of the song entered the public domain in the Uni...

This article needs additional citations for verification. Please help improve this article by adding citations to reliable sources. Unsourced material may be challenged and removed.Find sources: Liberal Democratic Congress – news · newspapers · books · scholar · JSTOR (December 2009) (Learn how and when to remove this template message) Political party in Poland Liberal Democratic Congress Kongres Liberalno DemokratycznyLeaderJanusz Lewandowski (fir...
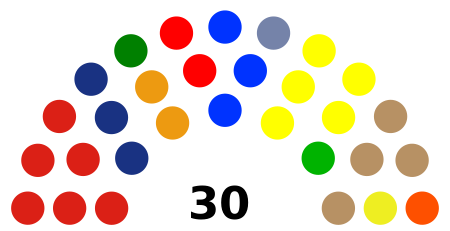

Dewan Perwakilan Rakyat Daerah Kabupaten Bengkulu UtaraDewan Perwakilan RakyatKabupaten Bengkulu Utara2019-2024JenisJenisUnikameral SejarahSesi baru dimulai9 September 2019PimpinanKetuaSonti Bakara, S.H. (PDI-P) sejak 18 Oktober 2019 Wakil Ketua IJuhaili, S.IP. (Golkar) sejak 18 Oktober 2019 Wakil Ketua IIHerliyanto Hazadin, S.IP. (Gerindra) sejak 18 Oktober 2019 KomposisiAnggota30Partai & kursi PDI-P (6) NasDem (3) PKB (1) Hanura...

Drama organization in New York City This article is about the professional association. For the television series, see Actors Studio (TV series). Actors StudioFacade in 2018FormationOctober 5, 1947; 76 years ago (October 5, 1947)TypeDrama schoolPurposeOrganization for professional actors, theatre directors and playwrightsHeadquarters432 West 44th StreetManhattan, New York CityRegion served United StatesKey peopleAl Pacino, Alec Baldwin, Ellen Burstyn (co-presidents)Beau Gravitte ...

Building to house a person who watches for wildfires A fire lookout tower on Vetter Mountain in Angeles National Forest, near Los Angeles, California Beyazıt Tower, an 85-meter-tall (279 ft) fire lookout tower at Beyazıt Square in Istanbul A fire lookout tower, fire tower, or lookout tower is a tower that provides housing and protection for a person known as a fire lookout, whose duty it is to search for wildfires in the wilderness. It is a small building, usually on the summit of a mo...

The Exorcism of Emily RoseLaura Linney e Tom Wilkinson in una scena del filmTitolo originaleThe Exorcism of Emily Rose Lingua originaleinglese Paese di produzioneStati Uniti d'America Anno2005 Durata119 min Rapporto2,35:1 Genereorrore RegiaScott Derrickson SceneggiaturaPaul Harris Boardman, Scott Derrickson ProduttorePaul Harris Boardman, Tom Rosenberg, Gary Lucchesi, Tripp Vinson, Beau Flynn Produttore esecutivoAndre Lamal, Terry McKay, David McIlvain, Julie Yorn Casa di produzioneScreen Gem...

هذه المقالة عن المعتقد المسيحي لعودة المسيح). لمعانٍ أخرى، طالع المجيء الثاني للمسيح (توضيح). المجيء الثاني للمسيح زجاج النافذة الملون في كنيسة سانت ماثيو الإنجيلية اللوثرية الألمانية في تشارليستون، جنوب كارولينا. في المسيحية والإسلام، المجيء الثاني للمسيح أو الظه�...

Historic oil storage tunnels in Darwin, AustraliaThis article needs additional citations for verification. Please help improve this article by adding citations to reliable sources. Unsourced material may be challenged and removed.Find sources: Darwin oil storage tunnels – news · newspapers · books · scholar · JSTOR (May 2022) (Learn how and when to remove this message) Darwin Oil Storage Tunnels in 2016 The Darwin oil storage tunnels were built during ...

Private university in Tokyo, Japan This article needs additional citations for verification. Please help improve this article by adding citations to reliable sources. Unsourced material may be challenged and removed.Find sources: Daito Bunka University – news · newspapers · books · scholar · JSTOR (February 2017) (Learn how and when to remove this message) Daito Bunka University大東文化大学Higashimatsuyama campusTypePrivate universityEstablished1...

Drum machine TR-505The Roland TR-505ManufacturerRoland CorporationDates1986–1991Price$318 US (1986)Technical specificationsPolyphony8 voicesOscillatorN/ASynthesis typeDigital sample-basedVelocity expressionNoStorage memoryPatterns: 48 user, 48 preset. 6 Songs.EffectsNoHardwareMain panel features a simple LCD display, 15 buttons, 2 knobs, 16 trigger pads, 2 outputs for left and right/mono, headphone jack, and tape input/output.Input/outputKeyboard16 pattern keysExternal controlMIDI in/out, s...

Australian Aboriginal sign language Warlpiri Sign LanguageRdaka-rdakaRegionNorth Central Desert, AustraliaNative speakersNoneLanguage familyPama–Nyungan Ngumpin–YapaNgarrkicWarlpiriWarlpiri Sign LanguageLanguage codesISO 639-3–GlottologNone Warlpiri Sign Language, also known as Rdaka-rdaka (lit. hand signs),[1] is a sign language used by the Warlpiri, an Aboriginal community in the central desert region of Australia. It is one of the most elaborate, and certainly the most studie...
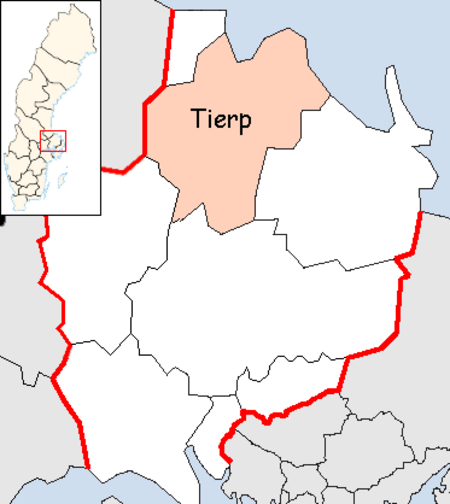
Artikel ini sebatang kara, artinya tidak ada artikel lain yang memiliki pranala balik ke halaman ini.Bantulah menambah pranala ke artikel ini dari artikel yang berhubungan atau coba peralatan pencari pranala.Tag ini diberikan pada Februari 2023. Munisipalitas Tierp atau Tierp Municipality (Tierps kommun) adalah sebuah kotamadya di Uppsala County pada tengah timur Swedia. Daerah ini terletak di kota Tierp. Referensi Pranala luar Tierp Municipality - Official site

У этого топонима есть и другие значения, см. Сыромятническая улица. Нижняя Сыромятническая улица Общая информация Страна Россия Город Москва Округ ЦАО Район Басманный Протяжённость 360 м Метро Курская Курская ЧкаловскаяD2D4 Курская (МЦД) Прежние названия Сыромятн�...

Cola di Rienzo Información personalNacimiento 1313 Roma (Estados Pontificios) Fallecimiento 8 de octubre de 1354jul. Roma (Estados Pontificios) Información profesionalOcupación Político [editar datos en Wikidata] Epistolario di Cola di Rienzo Nicola Gabrini, (* Roma 1313 – † 1354), más conocido como Cola di Rienzi y Cola di Rienzo, fue un notario papal y tribuno del pueblo romano y proclamó en Roma una nueva forma de gobierno inspirada en la República romana a la que llam...

LA Women's Tennis Championships 2009Sport Tennis Data3 agosto – 9 agosto Edizione36a SuperficieCemento CampioniSingolare Flavia Pennetta Doppio Chia-Jung Chuang / Zi Yan Il LA Women's Tennis Championships 2009 è stato un torneo di tennis giocato sul cemento. È stata la 36ª edizione del LA Women's Tennis Championships, che fa parte della categoria Premier nell'ambito del WTA Tour 2009. Si è giocato al Home Depot Center di Carson, vicino a Los Angeles negli Stati Uniti, dal 2 al 9 agosto ...

Rock Cup 2013-2014 Competizione Rock Cup Sport Calcio Edizione 60ª (1ª riconosciuta dalla UEFA) Organizzatore GFA Date dall'8 gennaio 2014al 10 maggio 2014 Luogo Gibilterra Partecipanti 21 Risultati Vincitore Lincoln Red Imps(15º titolo) Secondo Europa FC Statistiche Incontri disputati 17 Gol segnati 85 (5 per incontro) Cronologia della competizione 2012-2013 2014-2015 Manuale La Rock Cup 2013-2014 è stata la 60ª edizione della coppa di Gibilterra, la prima ri...

Tetangi MatapoTetangi Matapo in 2016Member of the Cook Islands Parliamentfor TamaruaIncumbentAssumed office 29 January 2013Preceded byPukeiti Pukeiti Personal detailsBorn3 October 1965Political partyCook Islands Democratic Party Tetangi Matapo (born 3 October 1965)[1] is a Cook Islands politician and member of the Cook Islands Parliament. She is a member of the Cook Islands Democratic Party. Matapo was born on Mangaia and educated at Mangaia School and Mangaia College.[1]...

For the most recent participation, see Italy in the Eurovision Song Contest 2024. Italy in the Eurovision Song Contest Participating broadcasterRadiotelevisione italiana (RAI)Participation summaryAppearances48First appearance1956Highest placement1st: 1964, 1990, 2021Host1965, 1991, 2022 Participation history 1956195719581959196019611962196319641965196619671968196919701971197219731974197519761977197819791980198119821983198419851986198719881989199019911992199319941995199619971998 –&...